One program with multiple in silico tools for approaching complex risk assessments using a weight of evidence approach.
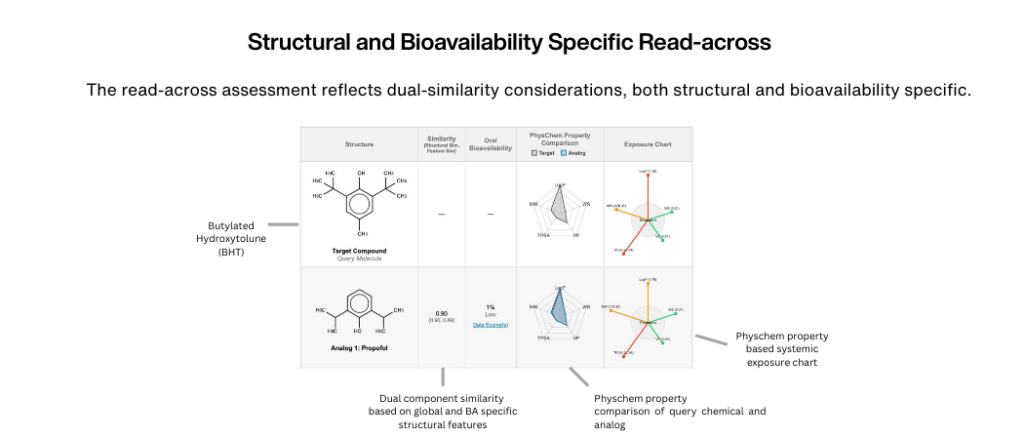
Using a multi-method approach, the QSAR Flex Bioavailability Module estimates the oral bioavailability of trace impurities and extractables and leachables, a key factor in toxicity evaluations for container systems and medical devices.
Oral Bioavailability Data: 1594 compounds.
Human Liver Microsomal Metabolism (HLM) Data: 4637 compounds.
CYP Substrate Datasets: CYP3A4 (2601), CYP2D6 (2249), CYP2C9 (2209)
MDR1 (P-gp) Substrate Data: 539 compounds


Values reflect the likelihood that the compound may undergo rapid metabolic degradation, primarily via Phase I oxoreductive pathways involving enzymes such as cytochrome P450s, flavin monooxygenases, esterases, and epoxide hydrolases.
Estimates the likelihood that the compound functions as a substrate for key solute carrier (SLC) and ATP-binding cassette (ABC) transporters, factors that shape its intestinal absorption, systemic distribution, and clearance.
Values represent the likelihood that the compound is a substrate for major human cytochrome P450 enzymes.
Assesses how different formulation vehicles may influence the compound’s oral bioavailability, based on its physicochemical properties and known formulation behavior patterns.
The FlexFilters platform enables the user to execute and integrate various computational toxicology tasks including read-across, molecular fragment handling, descriptor calculations, QSAR modeling, and predictions.
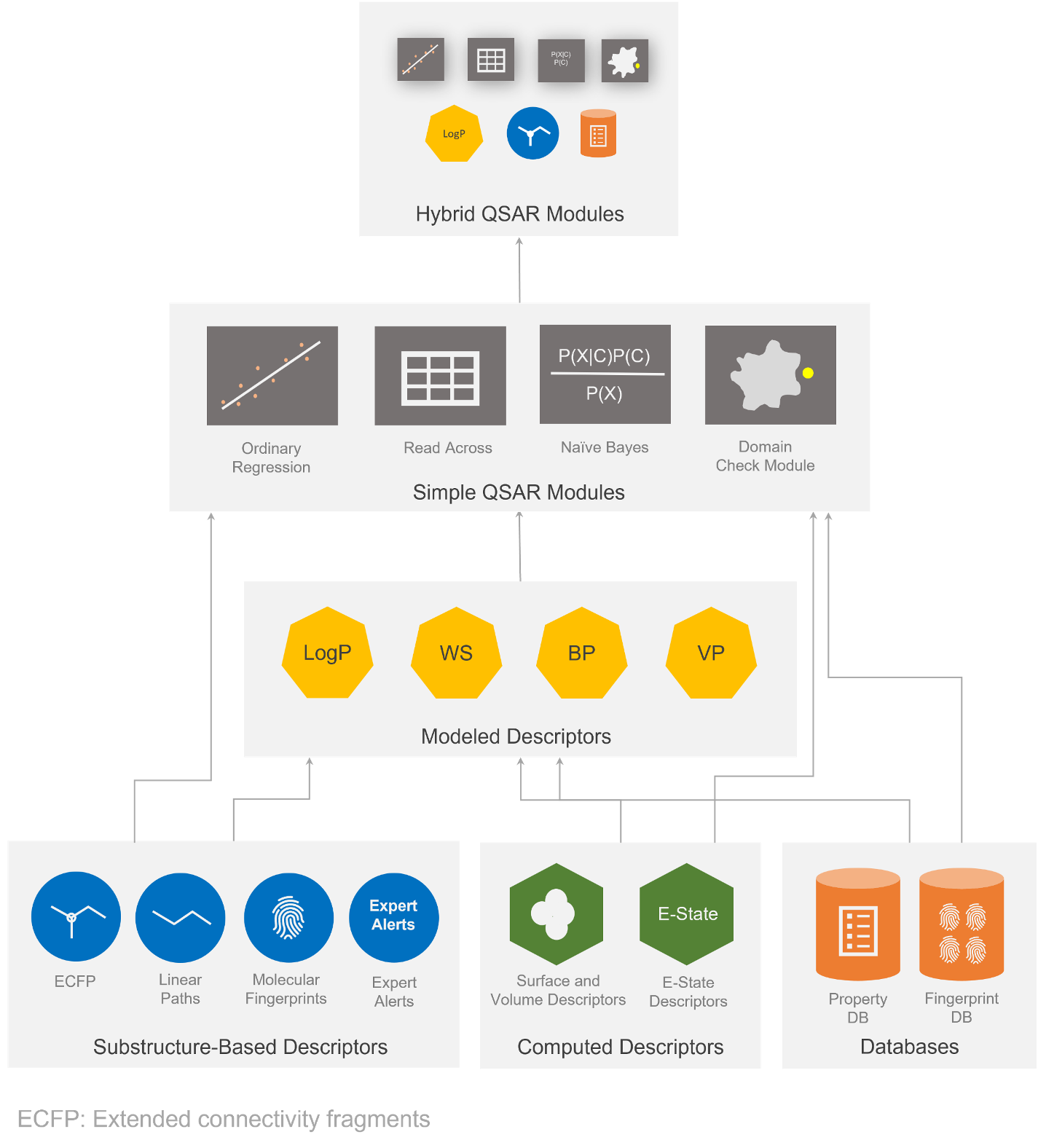
Evaluate the following endpoints with QSAR Flex:
Oral Bioavailability Assessment for Toxicity Evaluation
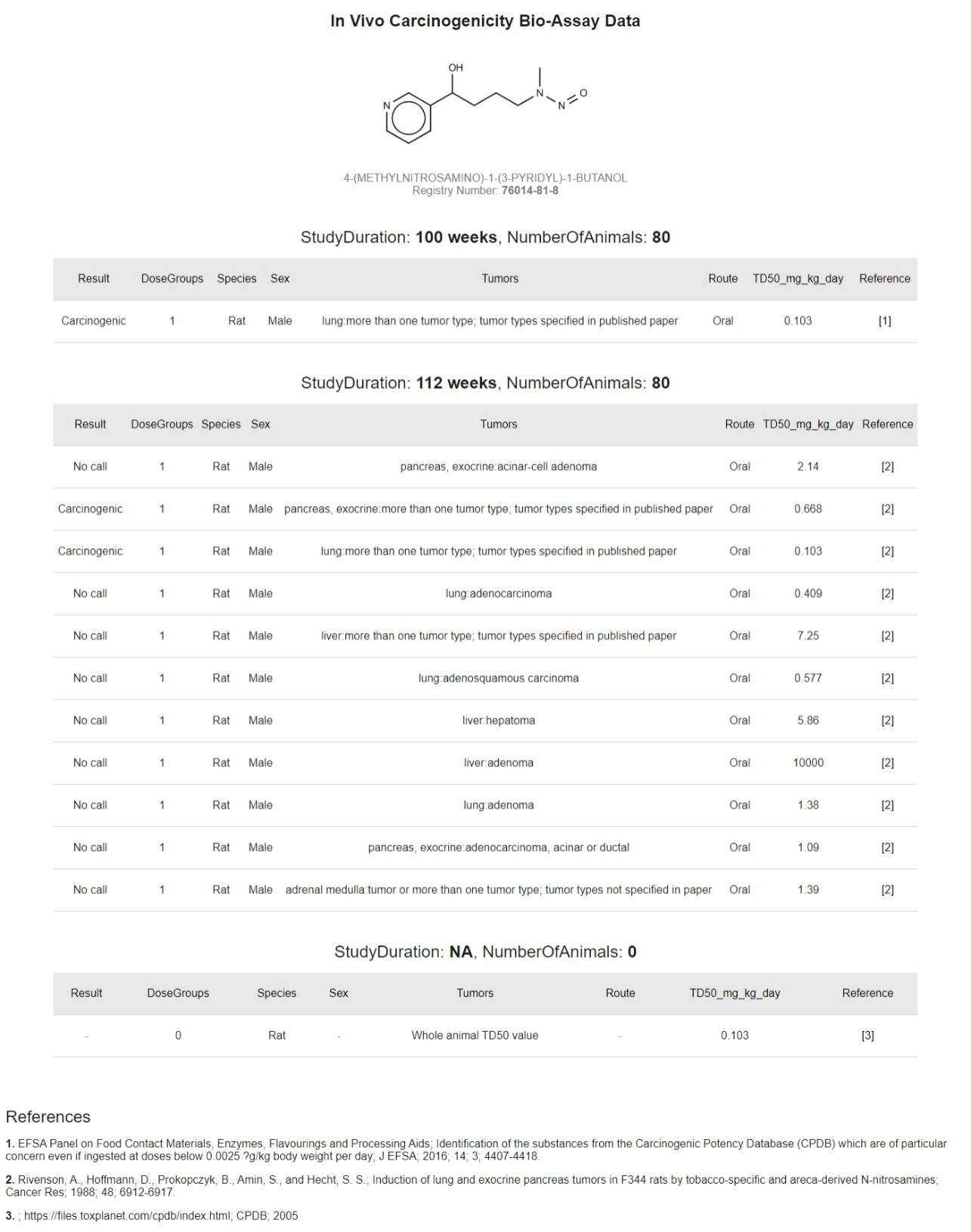
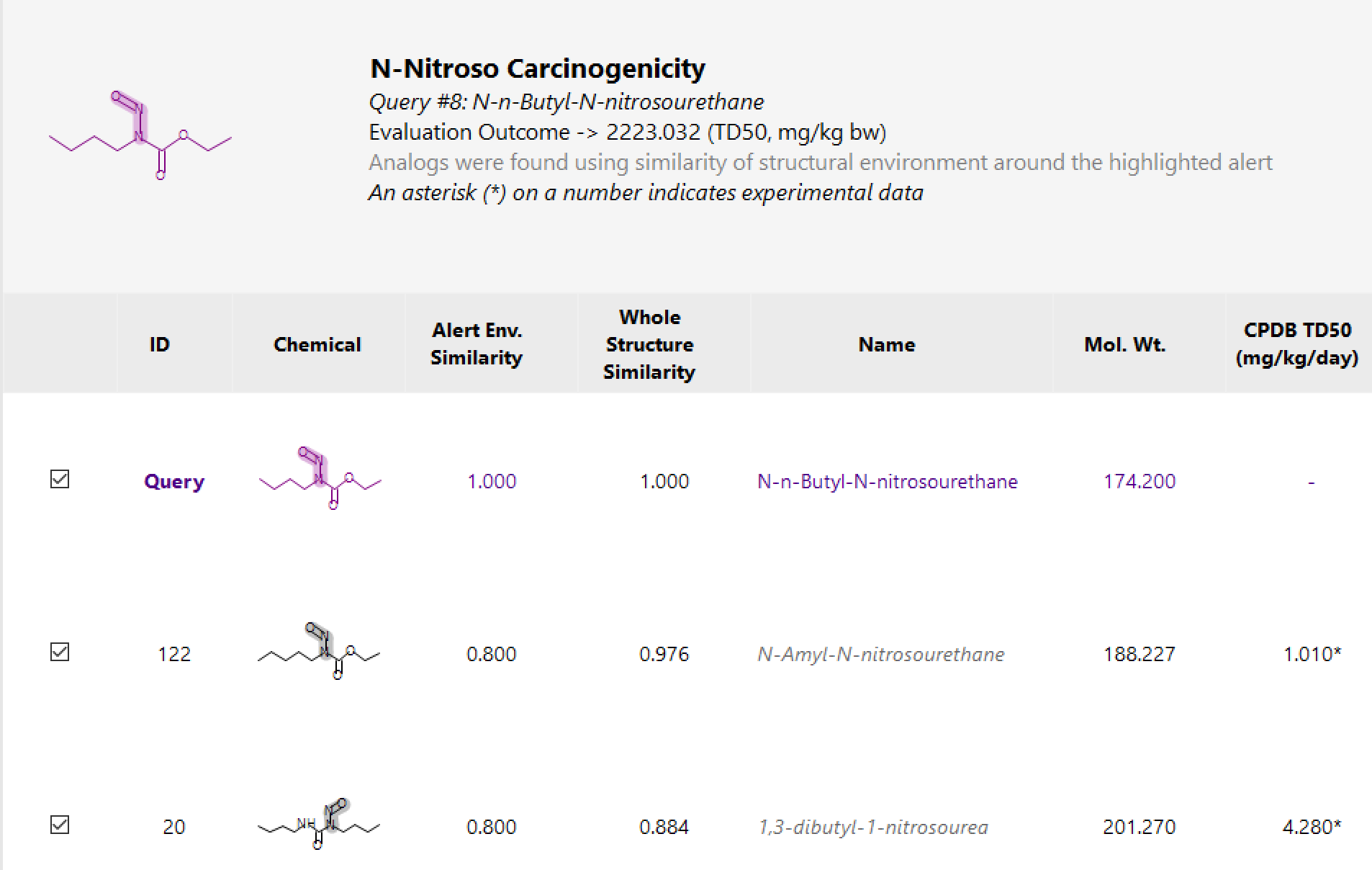
Carcinogenic Potency of N-Nitrosamines more
Molecular Properties
Ecotoxicity
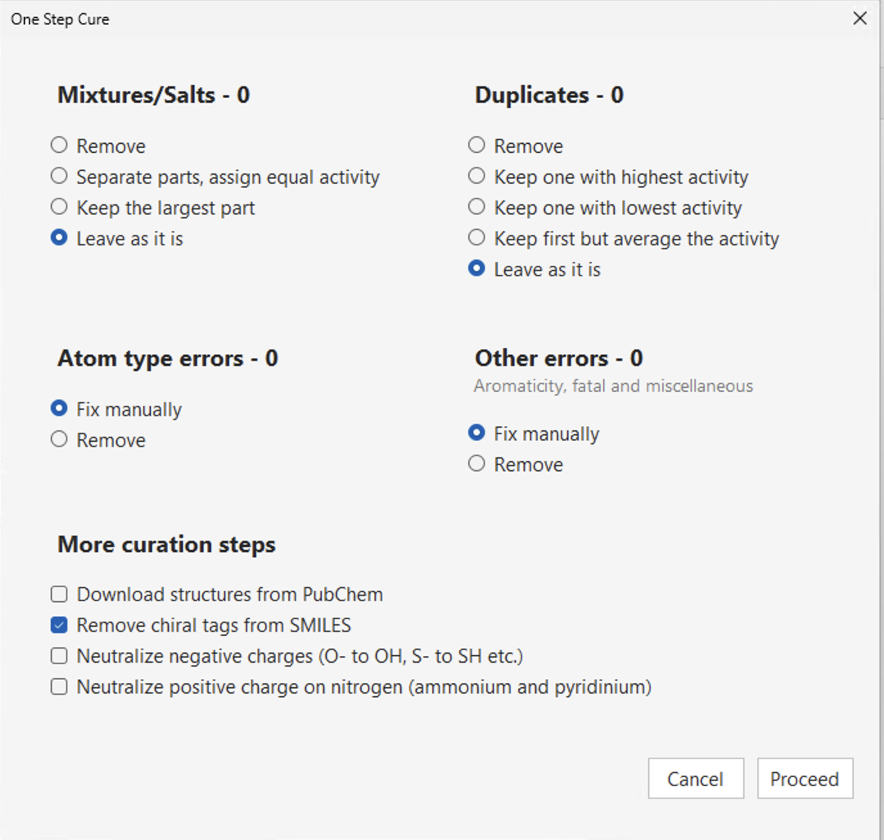
Curation of chemical data is a crucial starting step in any workflow and is carried out in QSAR Flex using the DataKurator tool.
License packages are customized to meet the needs of the user. Factors that affect pricing include length of license, number of endpoints, and number of users. Please contact us to request a quote.
Search for surrogates based on whole structure similarity or localized, fragment-based similarity.
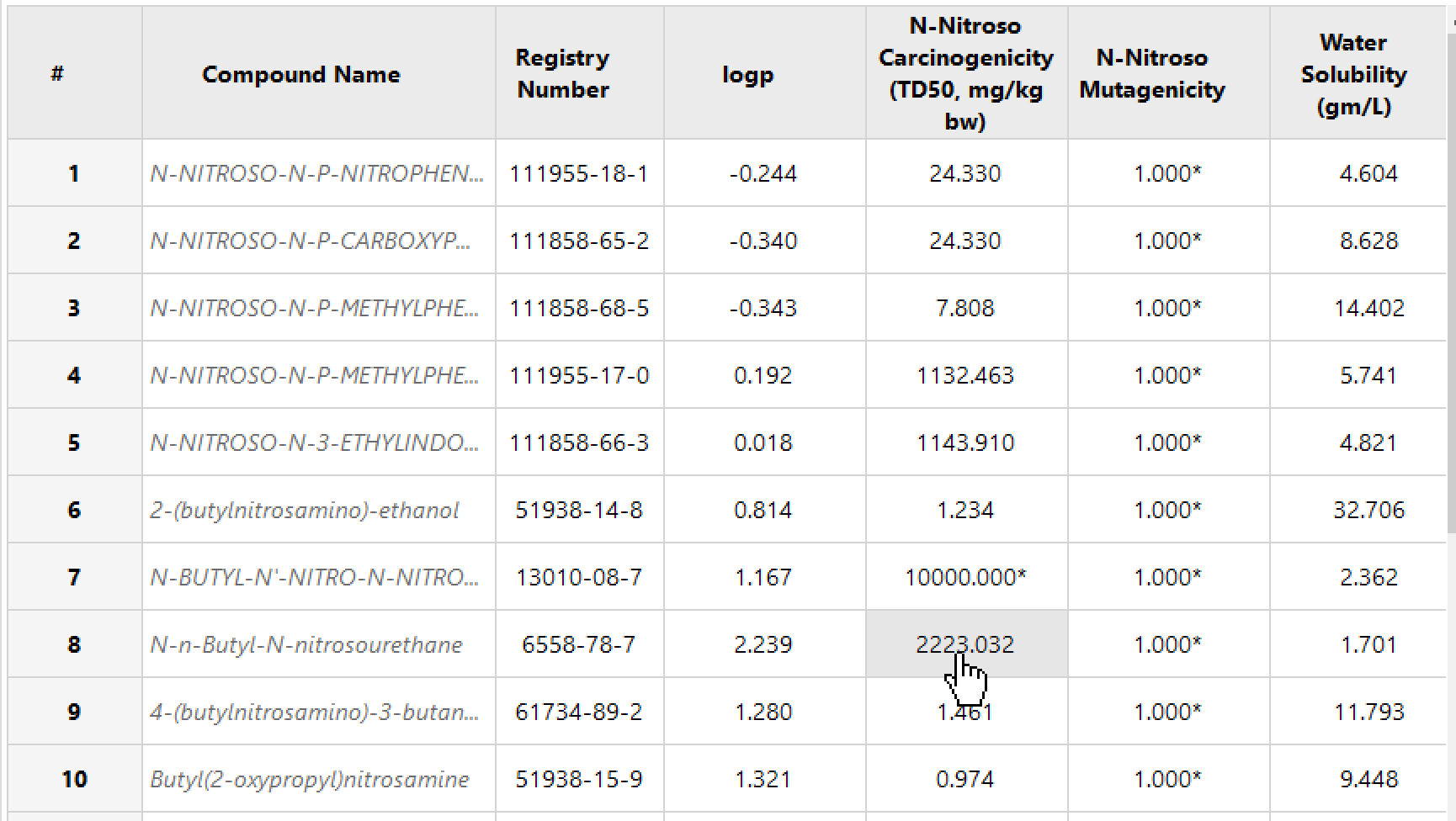
Batch processing for analog-based assessments.
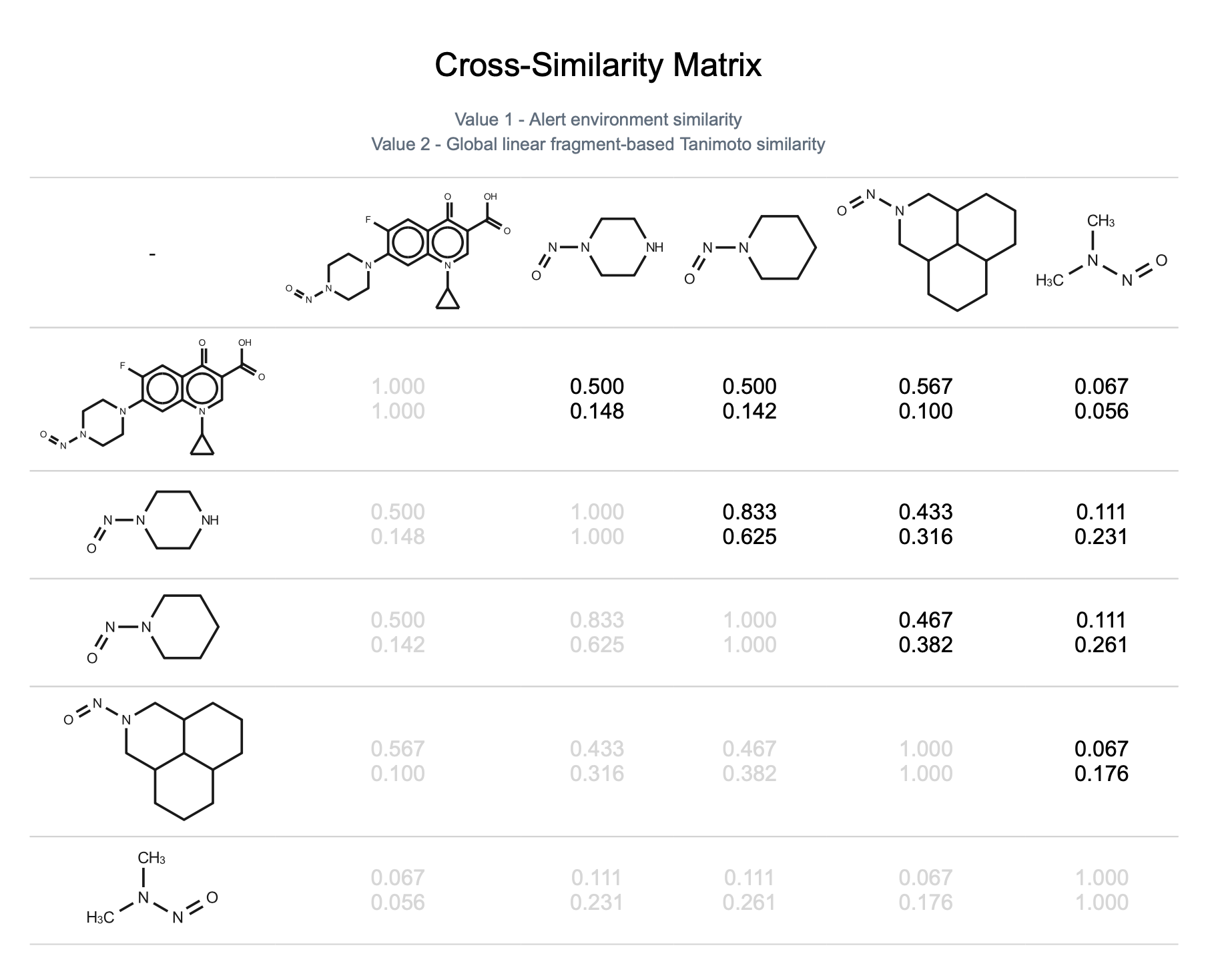
Analyze surrogate similarity using the Cross-Similarity Matrix.
Investigate experimental data of surrogates.